GEMV²
GEMV² is a Geometry-based Efficient propagation Model for V2V communication.
The original GEMV²¶
GEMV² has been developed by Mate Boban while working on his PhD thesis. There are three chief sources of information about this model:
- The GEMV² website: You can download the Matlab implementation and the user manual of GEMV² from there.
- Studying the code of this Matlab reference implementation
-
An article explaining some concepts behind the model:
Mate Boban, Joao Barros, and Ozan K. Tonguz: "Geometry-Based Vehicle-to-Vehicle Channel Modeling for Large-Scale Simulation", IEEE Transactions on Vehicular Technology, Volume 63, Number 9, November 2014 (DOI: 10.1109/TVT.2014.2317803)
A copy of this article is also available on the website.
GEMV² in Artery¶
This C++ implementation of GEMV² has been written from scratch by Thiago C. Vieira (Universidade Federal do Parana - UFPR) and Raphael Riebl (Technische Hochschule Ingolstadt - THI). However, Mate Boban's Matlab code and Artery's C++ version share some ideas, such as the usage of R-trees. While the Matlab implementation computes received power for each communication pair per time step at once, we compute the signal attenuation per transmission by implementing INET's IPathLoss interface. Hence, you can only use GEMV² with the INET radio model at the moment.
Action!¶
The animations below show the bundled visualizer module, which draws the outlines of vehicles (blue), buildings (black) and foliage (green).
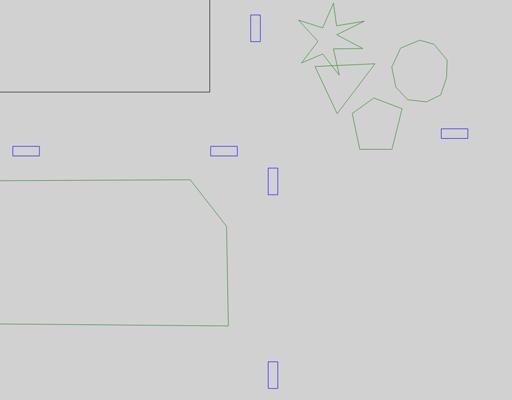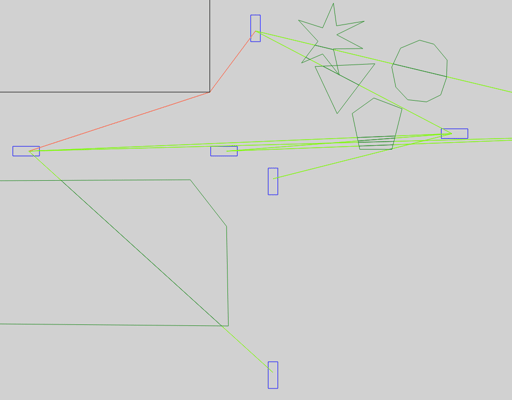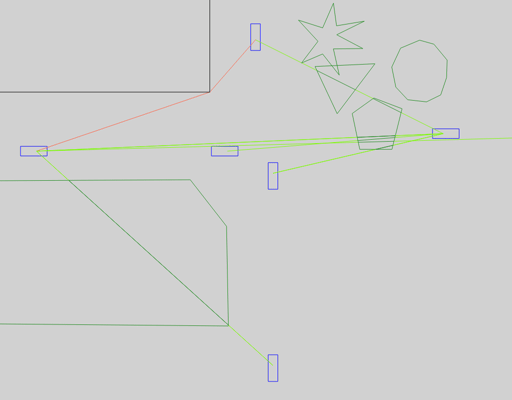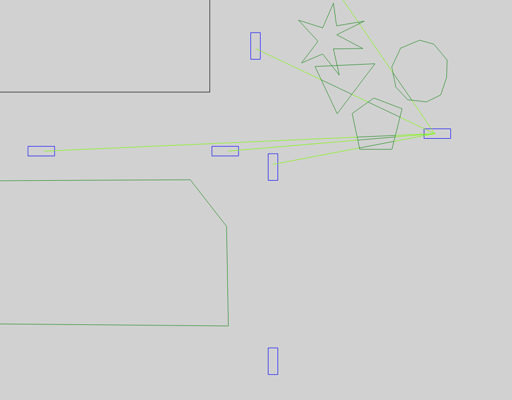
If GEMV² (artery.inet.gemv2.LinkClassifier) classifies a communication link to be mainly affected by buildings, it passes further path loss calculations on to its NLOSb model.
This model considers diffractions (orange) at the corners of buildings and onefold reflections (purple).

Foliage (single trees or forests) contributes attenuation based on total length a ray passes through it.
Our artery.inet.gemv2.FoliageIndex loads those polygons matching its filterTypes parameter from SUMO.
These polygons are treated as foliage by the NLOSf model then.
Artery's implementation supports concave, convex and overlapping foliage.